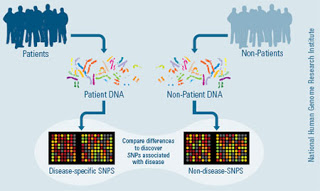
A recent article just published in Nature Genetics by Kottgen et al identifies several loci which appear to confer susceptibility to the development of CKD. How did researchers identify these genes? By a genome-wide association study. Briefly, researchers obtained genetic samples from some of the largest existing public health databases–such as the Framingham Heart Study, just to name one–and performed a genome-wide associated study in order to identify single-nucleotide polymorphisms (SNPs) that are associated with a low estimated GFR (using either creatinine or cystatin as the measurement of GFR). They identified significant SNP associations with low GFR in the genes UMOD, SHROOM3, GATM-SPATA5L1, CST, and STC1.
Of these genes, perhaps the most convincing association was seen with UMOD, the only one of these genes which was associated with an estimated GFR low enough to be categorized as CKD, which they defined as Tamm-Horsfall protein, which is the most abundant protein in normal urine. Interestingly, mutations in UMOD do account for some rare forms of kidney disease, including juvenile hyperuricemic nephropathy, glomerulocystic kidney disease, and medullary cystic kidney disease type 2. Also, knockout of the UMOD gene in mice is associated with a decrease in their eGFR. UMOD gene expression occurs primarily in the thick ascending limb of the loop of Henle, and therefore these new data imply that perhaps some of the pathogenesis of CKD dervies from the loop of Henle–and perhaps not all the action is in the glomeruli as has long been the focus. The UMOD story for CKD–as well as the other genes identified in this major association study–may be an interesting one to follow in the years to come.

