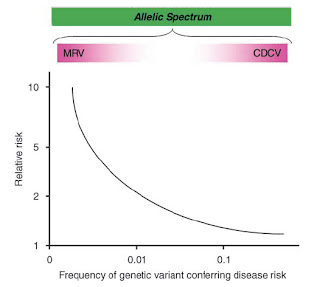
A paradigm of modern genetic studies, such as GWAS, is that there is a natural balance to allelic variation, with common variation in the population conferring mild disease risk and, conversely, genes of strong disease effect being rare. This phenomenon is formally described by the ‘common disease/ common variant’ (CDCV) and ‘multiple rare variant’ (MRV) models. The CDCV model predicts the existence of common genetic variants (present in 5–50%) that confer low to modest disease risk (e.g relative risks of 1.1–1.5). Complementing this is the MRV model, which holds that complex traits result from many different mutations, each of which is individually rare (a few percent at most and perhaps orders of magnitude less common), but with very strong effect (for example, relative risks 5 – 10, or even more; see figure). This phenomenon is thought to explain why the search for causative genes derived from GWA studies has been relatively unsuccessful, where only a handful of causative genes have been discovered in follow-up sequencing studies. It is assumed that this difficulty in finding culprit genes is due to these modest effects making them difficult to recognize. Undiscovered common genes of strong effect are simply not thought to exist…
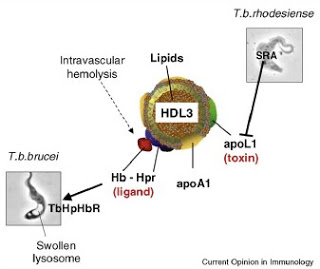
So what is going on in Nephrology? Within the space of a few short weeks there have been 2 separate, high quality studies identifying common disease-causing gene polymorphisms of very strong effect. First, there was the identification of 2 independent APOL1 variants, present in over 30% of African-American chromosomes, that carry odds ratios of 10.5 in idiopathic FSGS and 7.3 in hypertension-attributed ESRD. Analagous to HgbS-mediated protection from malaria, these APOL1 risk variants appear to have risen to very high frequency in Africa as they cause resistance to trypanosomal infection, thus protecting from sleeping sickness. Although the mechanism by which APOL1 variants cause kidney disease are not known, the mechanism of trypanosomal resistance has been described, and offers the hope for a new treatment of this life-threatening infection.
And now this morning, investigators from Hong Kong, reporting in JAMA, describe finding another set of common gene polymorphisms of strong effect. This candidate gene study of Chinese patients with type 2 diabetes, identified several variants of the PRKCß 1 gene as being associated with incident ESRD and, in a follow-up analysis, CKD. The adjusted risk for ESRD was 6.04 (95% CI, 2.00-18.31) for individuals with 4 risk alleles compared with those with 0 or 1, and allele frequencies were high (7-12.2%). PRKCß 1 is an excellent candidate for kidney disease risk in diabetes, with a prior RCT of blockade of its gene-product, PKC-ß, demonstrating slowed disease progression and reduced proteinuria in diabetic patients already on maximal medical therapy.
So, recent studies in Nephrology are bucking the trend in genetic epidemiology, and challenging one of its most basic hypotheses. Whether this is because the assumptions themselves are flawed, or that Nephrology research is just catching up with other fields, remains to be seen.